AnVIL Community Conference 2025 Conference Report
September 03 - 05, 2025 Vanderbilt University Medical Center, Nashville, Tennessee
The 2025 AnVIL Community Conference (ACC25) brought together AnVIL program leads, developers, researchers, students, and community members for the second year of this growing event. The meeting featured invited presentations showcasing research conducted on AnVIL, a hands-on Collaboration Fest (CoFest!) focused on analysis topics spanning polygenic risk scores to reproducibility on AnVIL, and a community town hall.

Participation grew significantly compared to 2024, reflecting expanding engagement with the platform. A total of 213 participants joined the conference:
- 83 in-person attendees, including 14 students and 35 AnVIL team members.
- 130 virtual attendees, including 28 international participants, 30 students and 9 AnVIL team members.
- 72 virtual workshop participants
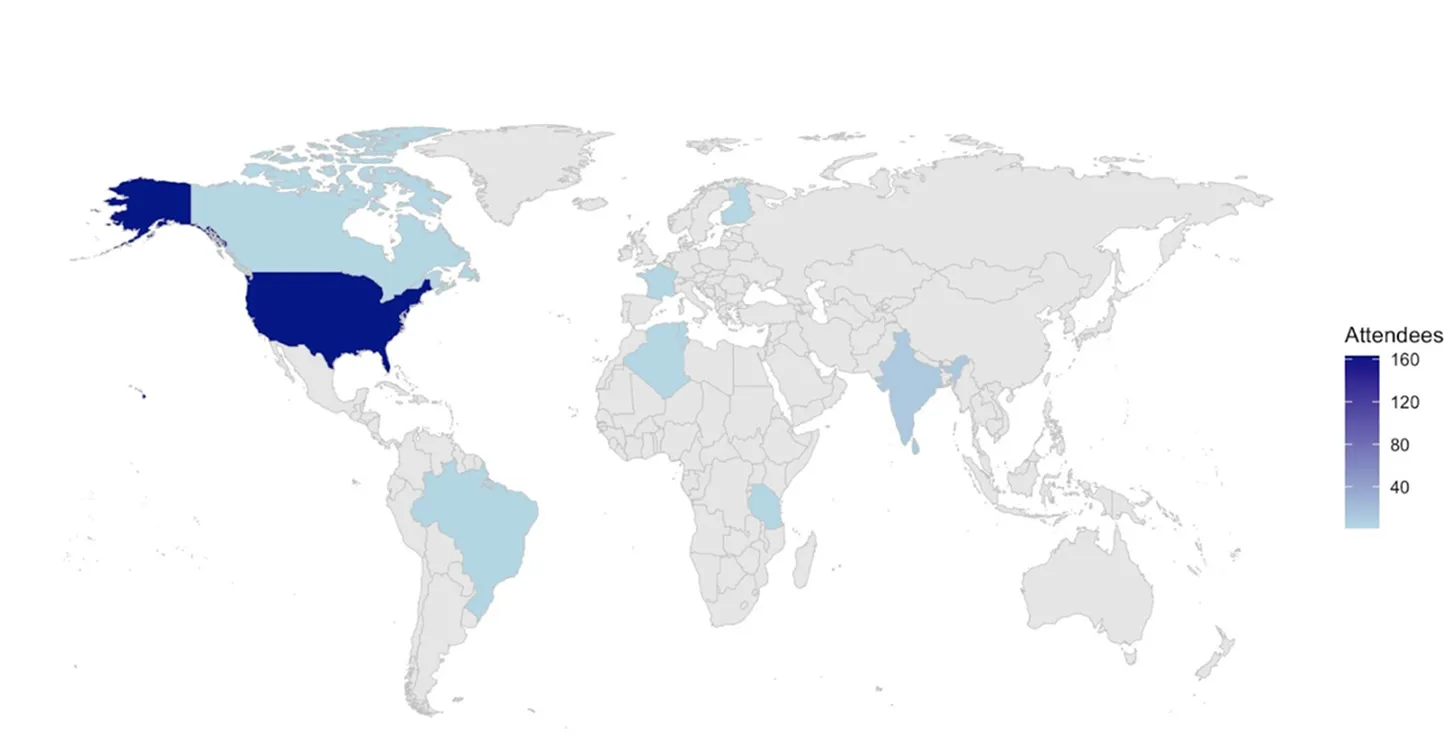
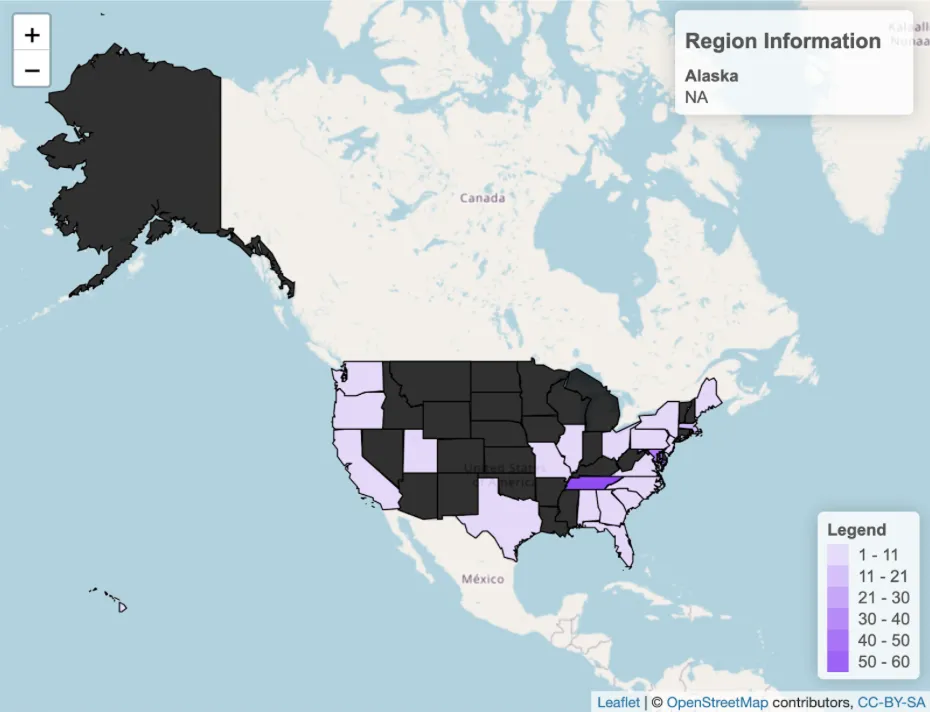
Attendees represented academic, government, and commercial organizations from across the United States and internationally, underscoring the increasing reach of AnVIL in the global genomics research community.
AnVIL Welcome and Roadmap
Robert Carroll, Vanderbilt University Medical Center, Nashville, TN
Jonathan Lawson, Broad Institute of MIT and Harvard, Cambridge, MA
Michael Schatz, Johns Hopkins University, Baltimore, MD
The second AnVIL Community Conference opened with a welcome from the three AnVIL program directors, led by Robert Carroll as host. Together they reflected on AnVIL’s growth and impact over the past year and presented the roadmap for the next phase of the project. Their talks showcased progress in clinical data resources, infrastructure for clinical data and analysis, and the integration of new tools and references to accelerate genomic discovery.
Clinical and Phenotypic Data – Robert Carroll
Robert Carroll welcomed attendees and highlighted AnVIL’s expanding role in supporting clinical and translational genomics. He described AnVIL's leadership in developing standardized data models, dictionaries, and provenance tracking, as well as modernized pipelines that make clinical data harmonization more efficient, reproducible, and scalable. Updates included progress on the i2b2 platform for federated data querying, as well as new pipelines for polygenic risk scores and kinship analysis. Advances in seqr were also discussed, including a scalable back-end, variant-level matching across ~65,000 exomes and genomes, and new workflows enabling patient-contributed data through GenomeConnect. Robert emphasized that these innovations are positioning AnVIL to support harmonized clinical data at scale, bringing together consortia, clinicians, and researchers in a secure environment.
Terra Updates – Jonathan Lawson
Jonathan Lawson shared updates from Terra, the cloud orchestrator within AnVIL that provisions secure resources and coordinates workflows across tools like Galaxy, Bioconductor, Dockstore, RStudio, and WDL pipelines. One significant announcement was a new imputation service jointly developed with the All of Us program. Built from a combined panel of genomes from both AnVIL and All of Us, this represents the largest imputation reference panel to date, with more than 515,000 genomes. Jonathan also described improvements to self-service data ingestion, which streamline data submission processes, and the introduction of quarterly data releases. AnVIL now hosts approximately 8.4 PB of data across more than 120 dbGaP accessions and over 100 consent codes. He previewed features from the Terra roadmap, including new cost management tools and expanded scientific services, emphasizing that the platform remains firmly focused on providing robust and sustainable access through Google Cloud.
Terra Feature Roadmap: https://support.terra.bio/hc/en-us/sections/30968105851931-Terra-Roadmap
The Next Phase of AnVIL – Michael Schatz
Michael Schatz discussed AnVIL’s ongoing integration of state-of-the-art genomic references and analysis pipelines. In addition to the well-used GRCh38 reference, recent accomplishments include onboarding the Human Pangenome Reference Consortium (HPRC), and the telomere-to-telomere (T2T-CHM13) reference. The AnVIL team is actively onboarding and assessing the best-in-class variant calling pipelines and tooling such as DeepVariant, DRAGEN, and Parabricks. Michael also highlighted advances in user support and experience: from Galaxy’s AI-driven analysis guidance to expanded Bioconductor resources and the deployment of local and commercial LLMs for interactive assistance. These capabilities aim to lower barriers for researchers of all backgrounds, ensuring they can harness the full potential of the AnVIL ecosystem for large-scale and AI/ML-driven genomics.
Together, the directors outlined a vision for AnVIL as a secure, interoperable, and researcher-friendly platform. The roadmap for the coming years emphasizes harmonized clinical and phenotypic data, robust Google Cloud infrastructure, and innovative tools that will empower the genomics community to pursue discoveries in precision health at unprecedented scale.
Invited Talks
The AnVIL Community Conference hosted four talks from investigator teams using the AnVIL and other cloud resources to support their genomic analyses.
Somatic loss of X and Y chromosomes affects tumor gene expression
Melissa A. Wilson, PhD, National Human Genome Research Institute
What’s New and What’s Next with the All of Us Research Program
Paul A. Harris, PhD, Vanderbilt University Medical Center
Ten years of genetic epidemiology in the cloud: How open architecture has won the day
Alisa Manning, PhD, Broad Institute of MIT and Harvard, Massachusetts General Hospital, Harvard Medical School
Our Dynamic Genome: how we leverage AnVIL to empower genomic discovery and clinical translation
Alexander G. Bick, MD, PhD, Vanderbilt University Medical Center
Submitted Talks
This year, the AnVIL Community Conference welcomed abstract submissions for abstract-selected talks from the developer and researcher communities using AnVIL.
The All of Us and AnVIL Imputation Service
Beth Sheets, MS, Broad Institute of MIT and Harvard
All of Us and AnVIL Imputation Service: https://terra.bio/introducing-the-all-of-us-anvil-imputation-service/
Decoding the Molecular Heterogeneity of Severe Acute Malnutrition through Integrated Multi-Omics and Cloud-Enabled, AI-Driven Analyses
Yixing Han, PhD, National Human Genome Research Institute
Exploring the genetic architecture of human social music-making enjoyment
Tanguy Rubat du Mérac, PhD Candidate, Vanderbilt Genetics Institute
AnVIL for education: Current directions and future challenges teaching undergraduate biology through course-based student research experiences on the cloud
Sayumi York, PhD, Notre Dame of Maryland University
Poster Session
Prior to the poster session, abstract submitters were invited to give Poster Lightning Talks to describe the work presented in their poster. The meeting featured 16 community-contributed posters.
Community Forum
The Community Forum was an open opportunity for a full audience discussion on any topics of interest to the community. The discussion spanned compliance with new federal data security guidelines, sustainability in community support, functionality to display summary-level results, engagement with a broader set of communities, and requirements of data access for potential researchers not connected to a dbGaP recognized institution.
The key takeaways from the community session were:
- AnVIL is compliant with the new NIH guidelines for secure storage of controlled access datasets (NOT-OD-24-157), meeting the NIST SP 800-171, and AnVIL serves as a good place for controlled access data storage. However, the administrative process with institutions and signing officials to trust & adopt AnVIL is not yet clear. There are educational and awareness needs to support institutions in embracing cloud-first and AnVIL.
- The AnVIL team also will be attending conferences like NCURA and ASHG to connect with and share information with administrative personnel on how to rely on AnVIL, as well as developing materials to support researchers to understand this.
- AnVIL’s userbase is growing and the team is eager to sustain user support for a growing community.
- The team relies on training trainers and building up community members to scale support and knowledge sharing among users. Over time, users posting their publications, publicizing their workflows through Dockstore and other resources, asking their questions on the Support Forum - all contribute to a rich resource ecosystem where users are able to learn from previous work, like we see in the Galaxy and Bioconductor communities.
- Developing Featured Workspaces as gold standard examples of data organization, analysis, and sharing will help contribute to this resource.
- Seeding the community with users who are able to engage deeply and encouraging them to use all the functions and features of AnVIL will grow up a community around these users’ or consortia’s resources, and highlight to other groups the advantages of using AnVIL.
- Users were interested in exploring more ways to share results and summarized results and statistics (e.g., beyond a GWAS). This functionality has been developed by groups in different ways, primarily the AnVIL team has seen users use the Workspace Dashboard space to share some results details or develop shiny apps that enable researchers to interact with an interface to explore data.
- There are a number of large biobanks that have done phenome or genome wide summaries, and users have a challenge interacting with the massive summary files. Each of these often comes with its own browser, but it’s challenging to look across these resources, and this could be an opportunity for AnVIL.
- AnVIL doesn’t currently support persistent user interfaces or multi-tenant apps, though these would enable some features folks are interested in.
- AnVIL holds legacy datasets from NHGRI consortia which are in process of going through harmonization to improve its utility and reusability.
- The community was interested to engage less traditional potential researchers on AnVIL (e.g., high school students, medical librarians, general public). The team has had focused efforts with the Genomic Data Science Community Network from a wide range of institutions R1s to Community Colleges and beyond. There have been efforts to engage with international communities as well with a virtual workshop in Sri Lanka in January 2025.
- A challenge is that cloud compute credits can be offered for a limited time, and it can be difficult to empower these groups when credits are used or the offering period expires.
- Another challenge is that users require backing by a recognized institution to be able to access controlled-access datasets. There are over 1.2 PB of non-restricted access datasets, which can be a very valuable resource for students & citizen scientists.
- AnVIL follows the mandate to protect participants’ data in accordance with their consent and aims to make data access and reuse more available to a broad set of researchers and potential researchers. The community is supportive of NIH and dbGaP, which manage these controls, to explore broadening methods of data access for students.
CoFests!
CollaborationFests!, also known as CoFests!, are collaborative work events where we expand and improve the AnVIL community and the AnVIL ecosystem. At ACC2025, there were three CoFest! Sessions.
Pre-conference Virtual AnVIL Workshop
Scientific Lead: Ava Hoffman, PhD, Fred Hutch Cancer Center
Outreach Coordinator: Natalie Kucher, Johns Hopkins University
Held ahead of the AnVIL Community Conference, this live, virtual workshop provided participants with an interactive introduction to the AnVIL platform. The two-hour session guided attendees through the fundamentals of navigating AnVIL, exploring data, and launching cloud-based analyses using state-of-the-art bioinformatics tools. Designed as an accessible entry point for new users, the workshop emphasized hands-on learning, allowing participants to familiarize themselves with the platform’s interface and core features before the in-person CoFest sessions.
The key takeaways from this session were:
- Provide context for people new to AnVIL before the conference and build excitement
- Increase accessibility for participants who can’t attend in-person
- Spread out time spent online to decrease Zoom fatigue
- Build awareness of training material and support form and grow community
AnVIL101
Scientific Lead: Ava Hoffman, PhD, Fred Hutch Cancer Center
Outreach Coordinator: Natalie Kucher, Johns Hopkins University
New to AnVIL? This hands-on CoFest! session provided participants with a comprehensive introduction to the platform. Attendees were guided through creating and managing an AnVIL workspace, accessing data from multiple sources including public repositories like the SRA, protected sources such as dbGaP, and personal datasets, and running analysis workflows in the cloud. The session emphasized practical exercises in moving data into Google Cloud, preparing Jupyter notebooks compute environments, and executing workflows to ensure reproducible analysis.
The key takeaways from this session were:
- Learned how to create and manage AnVIL workspaces and launch compute environments.
- Explored interactive compute environments via Jupyter notebooks to work with data.
- Gained hands-on experience importing data from SRA and personal sources into AnVIL.
- Learned about next steps for getting protected data (e.g., from dbGaP) onto AnVIL or locating data in the AnVIL Data Explorer.
- Discovered workflows using Dockstore.
- Ran workflows to transfer data using WDL scripts and cloud storage best practices.
- Developed familiarity with Google Cloud Storage concepts, including buckets and persistent disks, to manage workspace data.
This session lowered the entry barrier for researchers new to cloud-based genomic analysis, giving them the tools to leverage AnVIL’s capabilities effectively. Many participants were curious about next steps.

Polygenic Risk Score (PRS) Analysis in AnVIL
Scientific Lead: Matthew Lebo, PhD & Shruti Parpattedar, MS, Harvard Medical School
Outreach Coordinator: Elizabeth Humphries, PhD, Fred Hutch Cancer Center
This track, run by the AnVIL Clinical Resource team, consisted of both an overview and a hands-on workshop to provide individuals with an understanding of polygenic scores and how to run and evaluate them in AnVIL. First, we level-set by providing an overview of the current state of polygenic analysis, with a focus on polygenic risk scores (PRS). Next, we jointly worked with participants to run PRS mix analyses in AnVIL using the WDL framework. These tasks increased in complexity in terms of analytical components of the workflow, with the goal of enabling users to run the WDL on their own. We also engaged with participants to get feedback and create user-friendly documents to enable processing of this workflow once published to the broader community.
The key takeaways from this session were:
- Learned about Polygenic Risk Scores (PRS) and Polygenic Score (PGS) Catalogue.
- Learned about PRS mix scores and how to calculate them.
- Learned how to run a Polygenic Risk Score (PRS) WDL workflow on AnVIL.
- Lots of tracking and troubleshooting.

AnVIL Mini Hack: Reproducibility in Action!
Scientific Lead: Allissa Dillman, BioDataSage, Montgomery College
Outreach Coordinator: Kate Isaac, Fred Hutch Cancer Center
Reproducibility can be deceptively challenging in computational genomics. In this hands-on mini-hackathon, participants explored the intricacies of reproducing figures from the MAGE RNA sequencing study using real omics data from the 1000 Genomes Project and MAGE. Teams were encouraged not only to recreate published figures but also to design new visualizations from these open datasets. The session emphasized that computational reproducibility requires creativity, problem-solving, and detective work beyond simply following protocols.
The key takeaways for this session were:
- Hands-on experience with cloud-based tools on AnVIL to explore and reproduce RNA-seq analyses.
- Gained familiarity with public datasets from the MAGE and 1000 Genomes datasets.
- Learned best practices for managing and organizing analysis in an AnVIL workspace.
- Developed problem-solving skills for interpreting published results and creating new visualizations from open data.

Summary
The second AnVIL Community Conference brought together the leads, developers, and users of AnVIL to showcase exciting research performed on the platform, highlight current capabilities, and gather community input to guide future offerings and adoption of cloud computing for genomics research. Building on the inaugural event, ACC2025's expanded programming included poster sessions, submitted community talks, and lightning talks, creating a richer, more interactive program for participants.
Participation grew significantly compared to 2024, reflecting expanding community engagement. A total of 213 participants joined the conference: 83 in-person and 130 virtual attendees. Through keynote talks, hands-on workshops and collaboration sessions, poster presentations, and interactive discussions, the conference highlighted opportunities, priorities, and future directions for AnVIL to support the genomics community at scale.

Feedback
A few quotes from our attendees:
“I was very skeptical before I attended but now I am very interested in using AnVIL. The conference was a success! Thank you.”
“I thought that all of the talks were very engaging, I liked the pace of the meeting and the mix of speakers/topics. I also felt like most speakers were very energetic and the energy was very positive.”
“I'm glad that I attended this meeting!”
“[I liked the] Formal and informal networking opportunities; loved the Community Forum ("townhall" session); liked the buffet style food which encouraged additional networking”
“Great talks and community!”